|
[GT] 단백질 디자인에 인공지능이 사용되다 | |||
| ‘네이처 컴퓨테이셔널 사이언스(Nature Computational Sc... |


  |
 |  |
 |
 |  |
 |
[Global Technology Breifings]
While artificial intelligence will make a substantial impact in automating consumer oriented and everyday business applications, its greatest contribution will be in areas where humans are unable to make progress because of complexity.
That’s why new results related to using AI in protein design are so exciting.
The journal Nature Computational Science just published the results of research at the University of Toronto into using an artificial intelligence system to create proteins not found in nature.
This AI system uses generative diffusion, the same technology behind popular image-creation platforms such as DALL-E and Midjourney.
The system promises to speed drug developtirely new therapeutic proteins more efficient and flexible. The model learns to generate “fully new” proteins, at a very high rate, starting from image representations.
And all the proteins it generates appear to be biophysically real, meaning they fold into configurations that enable them to carry out specific functions within cells.
Proteins are made from chains of amino acids that fold into three-dimensional shapes, which in turn dictate protein function.
With a better understanding of how existing proteins fold, researchers have begun to design folding patterns not produced in nature.
But a major challenge has been to imagine folds that are both possible and functional. It’s been very hard to predict which folds will be real and work in a protein structure.
By combining biophysics-based representations of protein structure with diffusion methods from the image generation space, the researchers have begun to address this problem.
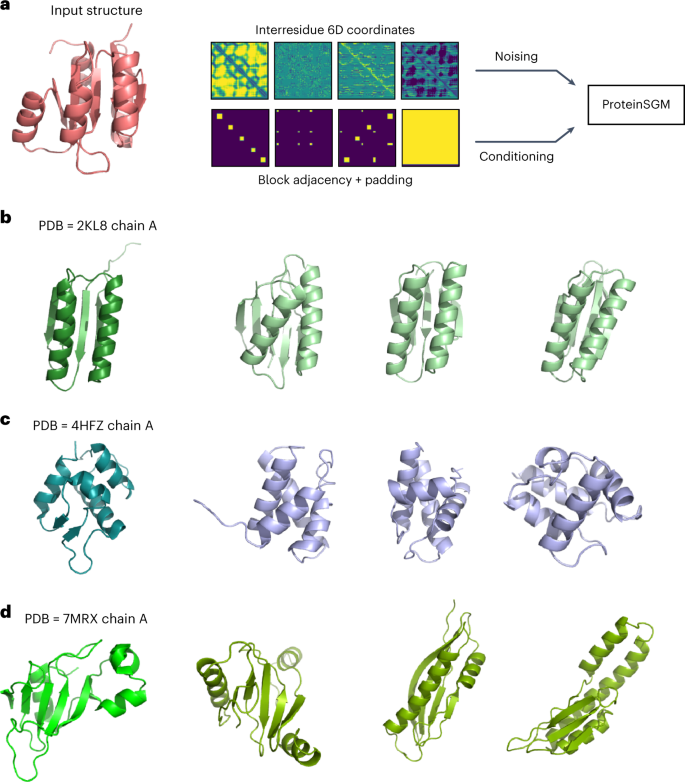
The new system, which the researchers call ProteinSGM, draws from a large set of image-like representations of existing proteins that encode their structure accurately.
The researchers feed these images into a generative diffusion model, which gradually adds noise until each image becomes all noise.
The model tracks how the images become noisier and then runs the process in reverse, learning how to transform random pixels into clear images that correspond to fully novel proteins.
To test their new proteins, the researchers first turned to OmegaFold, an improved version of DeepMind’s software AlphaFold 2.
Both platforms use AI to predict the structure of proteins based on amino acid sequences.
With OmegaFold, the team confirmed that almost all their novel sequences fold into the desired protein structures.
They then chose a smaller number to create physically in test tubes, to confirm the structures were proteins and not just stray strings of chemical compounds.
With matches in OmegaFold and experimental testing in the lab, they could be confident these were properly folded proteins.
They were amazed to see validation of these fully new protein folds that don’t exist anywhere in nature.
Next steps based on this work include further development of ProteinSGM for antibodies and other proteins with the most therapeutic potential.
This will be a very exciting area for research and entrepreneurship.
_아침독서상단(1).jpg)
.jpg)
.jpg)
.jpg)